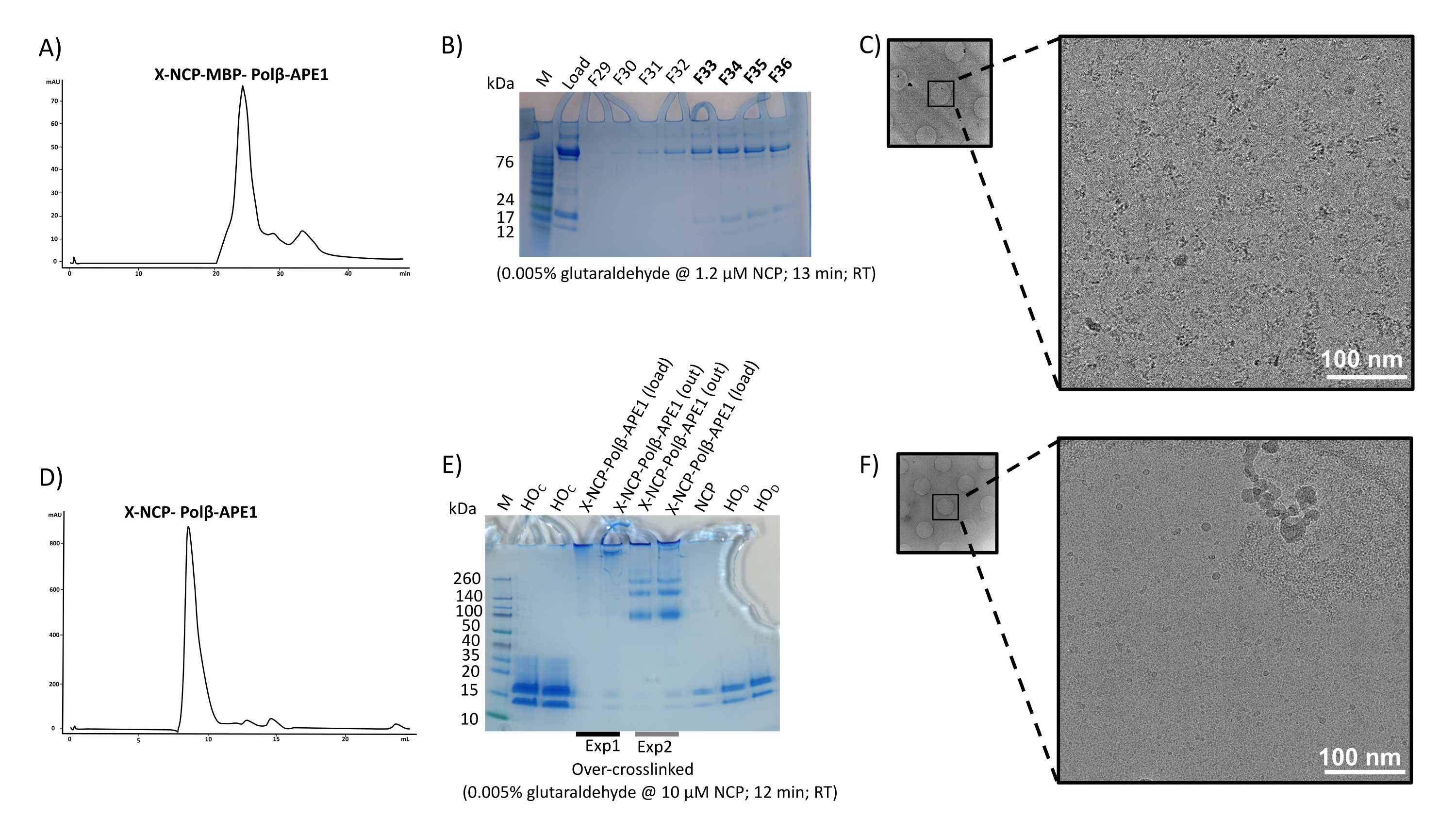
Preparation of Nucleosome Core Particles Complexed with DNA Repair Factors for Cryo-Electron Microscopy Structural Determination
Scientific Article | Ce protocole décrit en détail la préparation des complexes nucléosomiques à l’aide de deux méthodes de préparation des échantillons
DNA repair in the context of chromatin is poorly understood. Biochemical studies using nucleosome core particles, the fundamental repeating unit of chromatin, show most DNA repair enzymes remove DNA damage at reduced rates as compared to free DNA.

The cryo‐EM structure of the CENP‐A nucleosome in complex with
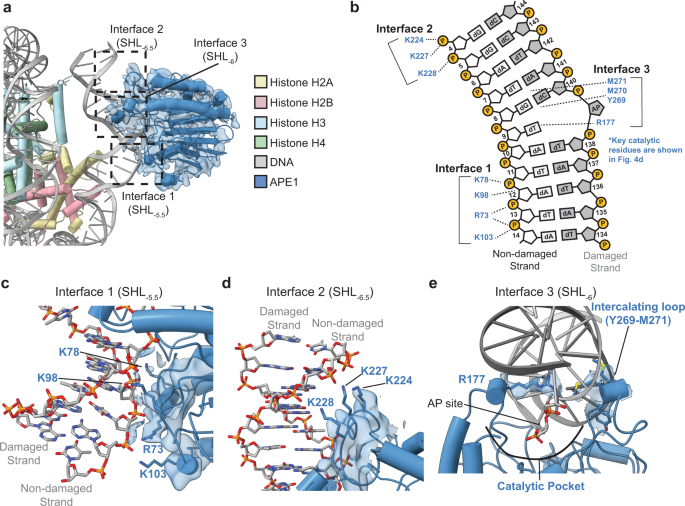
Structural basis for APE1 processing DNA damage in the nucleosome

Structural basis for APE1 processing DNA damage in the nucleosome

Cryo-EM structure of the human Sirtuin 6–nucleosome complex

Nucleosome Structure and Function

Self-Assembly of Geometry-Based DNA Origami-Histone Protein Hybrid
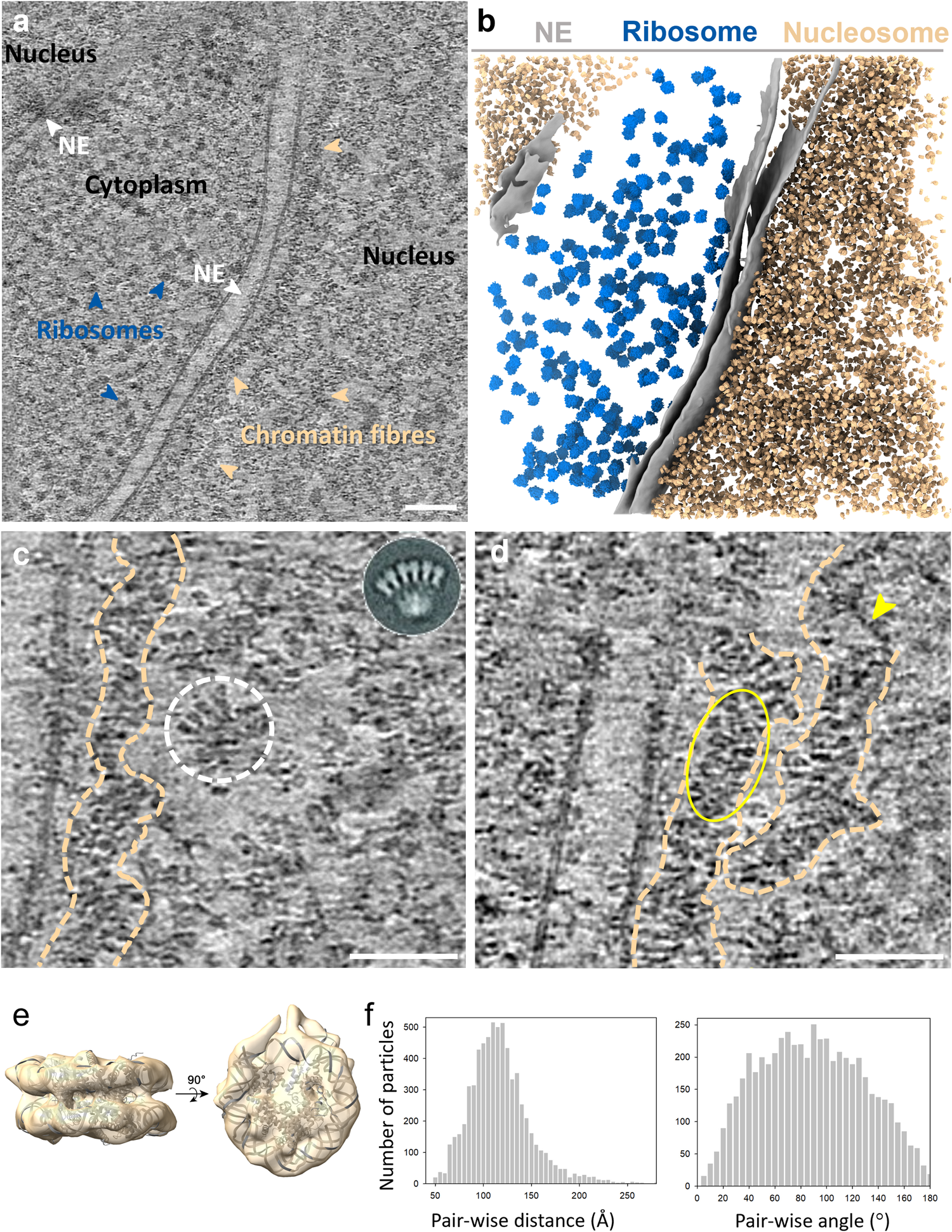
Structure of native chromatin fibres revealed by Cryo-ET in situ

Model for modulation of Polymerase enzymatic activities by the

Cryo-EM structure of a DNA-PK trimer: higher order oligomerisation

3.9 ˚ A phase-plate cryo-EM structure of the Nucleosome Core

Structures of +1 nucleosome–bound PIC-Mediator complex

Cryo-ET study from in vitro to in vivo revealed a general folding

Structural and mechanistic insights into the DNA glycosylase AAG

Cryo-electron microscopy structure of the H3-H4 octasome without

Cryo-EM structure of the nucleosome containing the ALB1 enhancer